
|
|
|
|
|
||
DOIs Available for Released Entries in the PDB ArchiveStructures released by the wwPDB into the PDB Archive are now being assigned a Document Object Identifier (DOI). The DOI System is used to identify content objects (such as journal articles, books, and figures) in the digital environment. The DOIs for PDB structures all have the same format – 10.2210/pdbXXXX/pdb – where XXXX should be replaced with the desired PDB ID. For example, the DOI for PDB entry 4HHB is "10.2210/pdb4hhb/pdb". This links directly to the entry in the PDB file format on the FTP server. The DOI can be used as part of a URL to obtain this data file (http://dx.doi.org/10.2210/pdb4hhb/pdb), or can be entered in a DOI resolver (such as http://www.crossref.org/) to automatically link to pdb4hhb.ent.Z on the main PDB ftp archive (ftp://ftp.rcsb.org). DOIs are automatically registered by the wwPDB when entries are released after the weekly update. They will not be available before a structure's release. Along with the ftp location, the DOIs for PDB entries also include the entry title, the authors, and the deposition date. RCSB PDB Focus: The ADIT Help SystemThe deposition tool ADIT (at RCSB or PDBj) includes examples and definitions provided in the PDB Exchange Dictionary as guides for users depositing their structures. An explanation for each piece of information requested by ADIT can be obtained by selecting the Help button located next to the named data item. This information will appear in the bottom frame. Pressing the Help button in the top frame will display these instructions. Examples for these items can be obtained by selecting the Example button within the table. This information will appear in the bottom frame. At any time during deposition, you may view the current state of the entire entry by pressing the PREVIEW ENTRY button. 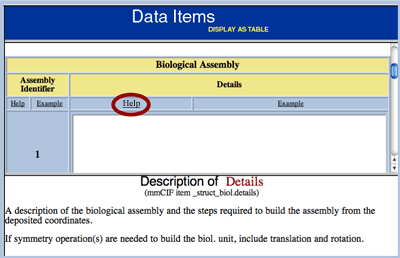 The ADIT help example for biological assembly details Questions about ADIT may be sent to deposit@deposit.rcsb.org. Annotation at the Research Collaboratory for Structural Bioinformatics Protein Data BankThe question "So, what does an 'annotator' do?" has been answered with the article:
RCSB PDB Focus: Depositing New Chemical Components (Ligands)
To deposit new ligands, please check Ligand Depot to see if the ligand, drug, ion, non-standard residue, modified residue, group, etc. is present in our chemical component dictionary.
If the ligand is not present in the dictionary, users can now upload a 2-D figure of the structure as part of the ADIT deposition process in PostScript, TIFF, or GIF formats. From the left-hand categories menu, scroll down and select the "Upload Supplemental Information: Ligand Information" to provide this information. Although the 3 letter code used for the ligand has no specific significance, you may check Ligand Depot and select a code for your ligand that has not been taken, otherwise, one will be selected for you.  Questions about ligands should be sent to deposit@deposit.rcsb.org. 2006 Deposition StatisticsIn 2006, 6911 experimentally-determined structures were deposited to the PDB archive. The entries were processed by wwPDB teams at the RCSB PDB, MSD-EBI, and PDBj. Of the structures deposited in 2006, 71.8% were deposited with a release status of "hold until publication"; 16.1% were released as soon as annotation of the entry was complete; and 12.1% were held until a particular date. 86.7% of these entries were determined by X-ray crystallographic methods; 12.8% were determined by NMR methods. 87.5% of these depositions were deposited with experimental data. During the same period of time, 6910 structures were released into the archive. |
||
©2007 RCSB PDB |