| OUTREACH AND EDUCATION
Explore a Structural View of Biology Using PDB-101

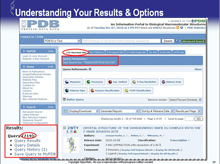
Click on the PDB-101 logo to access the education view; click on the blue logo in the top left at any time to access the main RCSB PDB website.
PDB-101 is a new and unique view of the RCSB PDB that places educational materials front and center. It packages together the resources of interest to teachers, students, and the general public to promote exploration in the world of proteins and nucleic acids.
Clicking on the blackboard PDB-101 logo (or its related widget in the left-hand menu) reveals the education-centered website. This view offers easy navigation: select any Molecule of the Month article from the top bar menu or mouse over the PDB-101 pull-down to jump to other sections of PDB-101.
This initial release of PDB-101 offers:
• Structural View of Biology. Built around the Molecule of the Month series, this feature promotes a top-down exploration of the PDB. Beginning with high-level functional categories, readers can browse through descriptive subcategories to access relevant articles that describe molecules in simple terms and access the related PDB entries. Mouseovers, pulldown menus, and carousels all offer easy navigational tools to promote learning.
• Molecule of the Month series. Since 2000, the RCSB PDB has published articles that describe the structure and function of a molecule along with interactive views, discussion topics, and links to structure examples. The collection of featured articles provides an annotated view of the PDB archive. With PDB-101, all Molecule of the Month issues appear on single pages, with links to printable PDF versions and downloadable high resolution images. They can be accessed through the pulldown menu in the top bar, the Structural View of Biology, and by archives organized by title, date, and category.
• Related Educational Resources and materials, including posters, animations, and classroom lessons and activities.
• Understanding PDB Data, a reference to help explore and interpret individual PDB entries. Broad topics include how to understand PDB data, how to visualize structures, how to read coordinate files, and potential challenges to exploring the archive.
PDB-101 will continue to be developed; we welcome your comments and suggestions.
To link directly to this view, use www.pdb.org/pdb-101.
Meetings and Events

At Experimental Biology, the RCSB PDB met with many users, including depositor Christopher Davies (Purdue University).

Molecule building at Rutgers Day

At ACA, the RCSB PDB Poster Prize was awarded to Briony Yorke for New Approaches to Time-Resolved Structural Studies of Macromolecules. Briony Yorke1, Arwen Pearson1, Mike Webb1, Robin Owen.2 (1The University of Leeds, Leeds, UK 2Diamond Light Source Ltd, Didcot, UK). Briony will receive a subscription to Science and a copy of The International Tables of Crystallography Volume F. Many thanks to the judges: Thomas Edwards (Emerald BioStructures), Katrina Forest (University of Wisconsin-Madison), and John Rose (The University of Georgia), and to Marcia Colquhoun and the ACA. The prize will also be awarded at the upcoming ISMB and IUCr
meetings.
At the Experimental Biology meeting (April 9-13, Washington DC), the RCSB PDB met with researchers and educators at the exhibit booth. Attendees, particularly from the American Society for Biochemistry and Molecular Biology, were interested in the searching and reporting features of the RCSB PDB website.
Rutgers Day, a full day of discovery and lively activities that showcase the varied resources, departments, and people at the university, was held April 30. The RCSB PDB was part of the Department of Chemistry & Chemical Biology's Bonding With Chemistry booth. Visitors of all ages built 3D DNA and virus structure models next to experiments and demonstrations of how chemistry impacts our lives.
The 2011 Meeting of the American Crystallographic Association (May 28-June 2, New Orleans, LA) was an active meeting. At the exhibition booth, visitors learned about new features such as the Validation Server, PDB-101, the wwPDB Common Tool for Deposition and Annotation, and much more. Director Helen Berman presented Putting the data in data mining: Curating the PDB archive as part of the Use of Databases in Structural Biology session. Lead annotator Jasmine Young described The wwPDB Common Annotation and Deposition Tool Development in a poster presentation, and the RCSB PDB Poster Prize for best student poster presentation involving macromolecular
crystallography was awarded.
Future meetings include:
• ISMB/ECCB: At the 19th Annual International Conference on Intelligent Systems for Molecular Biology and 10th European Conference on Computational Biology (July 15-19; Vienna, Austria), Senior Scientist Andreas Prlić will describe A Census of Internal Pseudo-Symmetries and Similarities in Protein Domains at the Laptop/Poster session at 3Dsig satellite meeting. Scientific Lead Peter Rose will help users Become an Expert User of the RCSB Protein Data Bank Website and Web Services at the Technology Track session on Monday, July 18 at 2:30 p.m.
• Protein Society: The PDB at 40: Past, Present, and Future will be discussed by Annotator Ezra Peisach during the poster presentations at this meeting (July 23-27; Boston, MA).
• IUCr: The XXII General Assembly and Congress of the International Union of Crystallography will be held August 22-30 in Madrid, Spain. Planned events include: a joint-wwPDB stand in the exhibition hall; a presentation on The wwPDB and Future Perspectives in Sharing Macromolecular Structure Data in the session on Developments and directions for crystallographic databases (Wednesday, August 24); an afternoon wwPDB Q&A session (Thursday, August 25); a discussion on Validation of small molecule and macromolecular X-ray structures. What are the differences and how can we learn from each other? by members of PDBe, CCDC, and RCSB PDB (Friday, August 26); and an introduction to The wwPDB Working Format: A Simplified Application of CIF Technology (Monday, August 29).
• PDB 40: Special symposium celebrating four decades of innovation in structural biology to be held October 28-30 at Cold Spring Harbor Laboratory. Early registration is strongly encouraged as the meeting is expected to fill quickly (meetings.cshl.edu/meetings/ pdb40.shtml).
New Publications
The RCSB Protein Data Bank: site functionality and bioinformatics use cases (2011) NCI-Nature Pathway Interaction Database Bioinformatics Primer doi: 10.1038/pid.2011.1
Miniseries: Illustrating the machinery of life: Eukaryotic cell panorama (2011) Biochem Mol Biol Educ. 39:91-101 doi:10.1002/bmb.20494
The evolution of the RCSB Protein Data Bank website (2011) WIREs Computational Molecular Science doi:10.1002/wcms.57
Narrated RCSB PDB Tutorial Updated
The narrated tutorial demonstrates and highlights how to use the tools found at www.pdb.org.
A comprehensive suite of RCSB PDB training materials is available at openhelix.com.
The updated training tools reflect many recent enhancements to the RCSB PDB site, including the data drill-down and data summary feature, updated ligand features such as a download page, images and binding affinity data, and new report types and visualization options.
The full tutorial runs for about an hour. Users can jump to any chapter: Introduction, Basic Searching & Browsing, Result Options, Structure Summary Page, Advanced Searching, Tools & Education, Summary, or Exercises.
The PowerPoint slides used as a basis for the tutorial, the suggested script for the slides, slide handouts, and exercises are all freely available for download and use as classroom content.
Congratulations to Science Olympiad Protein Modeling Champions

The first place team from Liberal Arts and Science Academy received their award from Fred Berry, the VP of Academics at the Milwaukee School of Engineering.
The National Science Olympiad Tournament was held May 20-21 at the University of Wisconsin at Madison.
Teams brought their A-game and some amazing prebuild models to the protein modeling competition.
The top scoring teams in this event were:
1) Liberal Arts and Science Academy (TX)
2) West Windsor-Plainsboro High School South (NJ)
3) Fairfax High School (VA)
Protein modeling will be offered as a 2012 event in the Science Olympiad; then it will be on hiatus from the tournament for two years as other events are incorporated. This event is managed by the MSOE Center for BioMolecular Modeling and hosted in NJ by the RCSB PDB. For protein modeling tips and news of interest to students and educators, follow us on twitter@buildmodels. |






