DATA
QUERY, REPORTING AND ACCESS
Website Statistics
Access statistics for the second quarter of 2011 are shown.
Month |
Unique Visitorss |
Visits |
Bandwidth |
April |
230825 |
557911 |
763.04 GB |
May |
233123 |
561880 |
814.08 GB |
June |
206456 |
507798 |
620.89 GB |
Enhanced Sequence Display
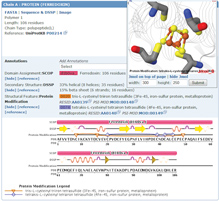
Protein modifications mapped onto the sequence diagram and Jmol view of ferredoxin (PDB entry 1b0v)
From each Structure Summary page, the sequence tab offers a diagram representation of the sequence. Each macromolecule chain can be annotated with domain assignments, secondary structure, and structural features such sites defined in the structure entry (i.e., binding sites of ligands) and protein modifications (i.e., posttranslational modifications). These annotations can be mapped on to the 3D view of the entry (jmol.sourceforge.net) for further exploration.
The current list of annotations includes:
• SCOP: domain annotations
• CATH: domain annotations
• Domain Parser (DP): domain annotations processed with the DP algorithm
• Protein Domain Parser (PDP): domain annotations processed with the PDP algorithm
• Pfam: regions with Pfam annotations
• Domain Parser (DP): domain annotations processed with the DP algorithm
• Protein Domain Parser (PDP): domain annotations processed with the PDP algorithm
• Pfam: regions with Pfam annotations
• Interpro: regions with Interpro annotations
• DSSP: secondary structure assignment
• STRIDE: secondary structure assignment
• Author Sec. Struc: secondary structure assignment as provided by
the entry's author
• NEW Protein Modification: protein modifications as detected
with software
• NEW Site Record: author assigned and software-detected binding sites
• NEW Single Nucleotide Polymorphism (SNP): data from the
LS-SNP database
Using the Display Parameters box, users can toggle between the display of unique chains and all chains. By default, only unique chains are displayed. This box can also toggle the display of the UniProtKB reference sequence.
Browse the PDB by GO Terms, EC Number, Source Organism, and More
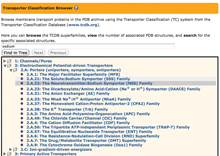
Use the Transporter Classification Browser to find PDB's membrane transport proteins as organized by TC Database family (www.tcdb.org).
The browser will autocomplete search terms with the matching classifications, and highlight locations on the tree.Use the Browse Database option to explore the PDB archive using different hierarchical trees. Browsers are available to search for related terms and structures based upon the following classifications:
• Trees for Biological Process, Cellular Component, and Molecular Function are organized using the Gene Ontology Consortium's descriptions for gene products (GO). PDB IDs and corresponding chain IDs have been mapped to GO terms by the SIFTS initiative.
• Enzyme Commission numbers. Search for enzymes by term or by partial or full EC number.
• Membrane Transport proteins organized using Transporter Classification (TC) Database system.
• Source Organism, using organisms found in the NCBI Taxonomy database. These organisms are the source of the individual naturally-occurring polypeptides. The PDB source organism assignment is based on author/UniProtKB-specified mapping of polypeptides.
• Genome Location of various organisms. The genomes represented are a subset of the genomes in the NCBI genome database and whose curated sequences for genetic loci are archived at Entrez Gene. The top level in the hierarchy
is the organism's genome. Each genome expands into chromosomes, which in turn expand into a list of loci on the chromosomes. Each locus is a link to retrieve structures associated with that locus.
• The MeSH terms used to classify publications indexed by the National Library of Medicine, and that appear in the entry's related PubMed abstract (Medical Subject Headings).
• SCOP description of evolutionary and functional relationships from the Structural Classification of Proteins.
• CATH clustering of proteins at 4 major levels from CATH: Protein Structure Classification.
The Browse Database feature can be accessed under the Search widget in the left-hand menu. Each tab links to a different browser.
Build Complex Queries with Advanced Search
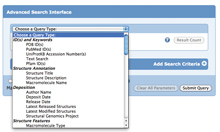
Combine different searches together to find structures and refine search results with Advanced Search.
New queryable options include protein modifications, Pfam ID, and EM structures with experimental data. Advanced Search provides the capability of combining multiple searches of specific types of data in a logical AND or OR. The result is a list of structures that comply with ALL or ANY of the search criteria, respectively.
Individual data items are organized by category; contextual help and examples are available by selecting the  icon. Recently added items include protein modifications, Pfam ID, and EM structures with experimental data. icon. Recently added items include protein modifications, Pfam ID, and EM structures with experimental data.
Currently, users can build searches based on:
• ID(s) and Keywords: PDB, PubMed, UniProtKB, Pfam IDs; text searching
• Structure Annotation: Structure title, description; and macromolecule name
• Deposition: Author name; Deposit Date; Release Date; Latest Released Structures; Latest Modified Structures; Structural Genomics Project
• Structure Features: Macromolecule Type; Number of Chains (asymmetric unit or biological assembly), entities; Models, and Disulfide Bonds; Molecular Weight; Secondary Structure Content; secondary structure length; SCOP, CATH, taxonomy
• Sequence Features: sequence; translated nucleotide sequence; sequence motif; chain length; protein modifications; genome location
• Chemical Components: name; ID; InChi descriptor; SMILES/SMARTS; molecular weight; chemical formula; binding affinity; has ligands; has modified residues
• Biology: Source; expression organism; Enzyme Classification; biological process; cell component; molecular function; Transporter Classification
• Methods: experimental method; X-ray resolution, R factor, diffraction source, reflections, cell dimensions, software, space group, crystal properties, detector; EM assembly
• Publication: citation; MeSH terms; PubMed abstract
• Misc: Has external links
The number of entries matching each individual query can be shown before running the full Advanced Search. Searches can be filtered by removing sequence similarity.
Queries built using Advanced Search can be stored in MyPDB to be run or modified at any time.
|



