DATA
DEPOSITION
2009 Statistics
In 2009, 8298 experimentally-determined structures were deposited to the PDB archive, and then processed by wwPDB teams at the RCSB PDB, PDBe, and PDBj.
Of the structures deposited, 74.6% were deposited with a release status of HPUB; 22.8% were released as soon as annotation of the entry was complete; and 2.6% were held until a particular date. 92% of these entries were determined by X-ray crystallographic methods; 7.1% were determined by NMR methods.
7448 structures were released in the PDB archive in 2009.
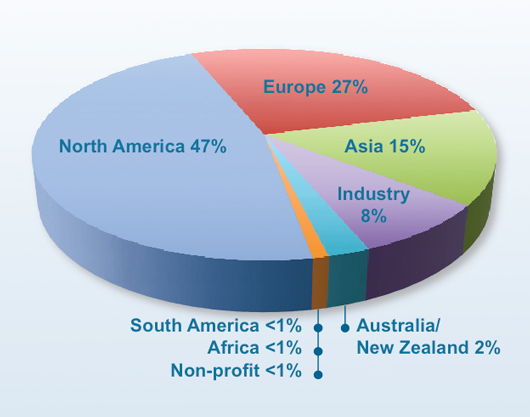
Depositor Locations (8298 structures deposited in 2009)
Data Deposition Resources
Planning to deposit a structure in 2010? There are several resources available to help.
The Validation & Deposition Portal contains links to software and documentation, including step-by-step deposition instructions specific for each experimental method: deposit.rcsb.org
The Deposition FAQ answers Frequently Asked Questions about depositing structures and releasing the processed PDB entries:
deposit.rcsb.org/depoinfo/depofaq.html
The Validation Server checks the format consistency of coordinate files and produces validation reports that can highlight potential problems before a structure is deposited: deposit.rcsb.org/validate/
For a depositor packet of printed materials, please send your postal address to to info@rcsb.org with the subject line first time depositor. |

