DATA
DEPOSITION AND ANNOTATION
2011 Deposition Statistics
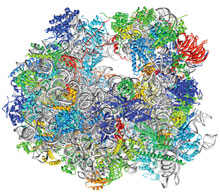 A record 30 ribosome structures were deposited in the fourth quarter of 2011, for a total of 49 ribosomes submitted in the year. These ribosomes include the crystal structure of the 80S ribosome from the yeast Saccharomyces cerevisiae shown here from PDB ID 3u5b.In the fourth quarter of 2011, 2295 experimentally-determined structures were deposited to the PDB archive for a total of 9250 entries deposited in the year. 8865 entries were deposited in 2010. A record 30 ribosome structures were deposited in the fourth quarter of 2011, for a total of 49 ribosomes submitted in the year. These ribosomes include the crystal structure of the 80S ribosome from the yeast Saccharomyces cerevisiae shown here from PDB ID 3u5b.In the fourth quarter of 2011, 2295 experimentally-determined structures were deposited to the PDB archive for a total of 9250 entries deposited in the year. 8865 entries were deposited in 2010.
Of all structures deposited in 2011, 81% were deposited with a release status of hold until publication; 17% were released as soon as annotation of the entry was complete; and 2% were held until a particular date.
93% of these entries were determined by X-ray crystallographic methods; 6% were determined by NMR methods.
8122 structures were released in the PDB archive in 2011. They account for 10% of the current total holdings of 78237.
Structural Genomics News
New Target Registration Database
 TargetTrack (sbkb.org/tt), a new experimental data tracking database, offers the latest information on the progress of structural studies on registered protein targets. TargetTrack (sbkb.org/tt), a new experimental data tracking database, offers the latest information on the progress of structural studies on registered protein targets.
TargetTrack provides information about protein complexes, membrane proteins, cryoEM studies, and other advances in structural biology along with the data previously available in TargetDB and PepcDB. Provided by worldwide structure genomics centers, these data can help many biological researchers with their projects' experimental design.
wwPDB News
X-ray Validation Task Force Report Published
To improve validation methods in the PDB, the wwPDB has convened method-specific Task Forces to collect recommendations and develop consensus on additional validation that should be performed, and to identify software applications to perform validation tasks.
The first report has been published by the X-ray Validation Task Force. These recommendations will be incorporated into the wwPDB data processing procedures and tools as part of the Common Deposition & Annotation Tool project.
A new generation of crystallographic validation tools for the Protein Data Bank. R.J. Read, P.D. Adams, W.B. Arendall III,
A.T. Brunger, P. Emsley, R.P. Joosten, G.J. Kleywegt, E.B. Krissinel,
T. Lutteke, Z. Otwinowski, A. Perrakis, J.S. Richardson, W.H. Sheffler, J.L. Smith, I.J. Tickle, G. Vriend, P.H. Zwart (2011) Structure 19:
1395-1412. doi:10.1016/j.str.2011.08.006
PDB Depositions Can Be Targets for CAPRI
 Depositing a protein-protein, protein-DNA, protein-RNA or protein-peptide complex? Consider submitting your structure as a
target for CAPRI after depositing your entry to the PDB. Depositing a protein-protein, protein-DNA, protein-RNA or protein-peptide complex? Consider submitting your structure as a
target for CAPRI after depositing your entry to the PDB.
CAPRI (Critical Assessment of PRedicted Interactions) is a community-wide, double-blind experiment aimed at assessing the performance of protein docking algorithms.
Submitting a target to CAPRI will help advance protein methods calculations, may provide new information on the quaternary structure of your complex, and should increase the visibility of your work. Moreover, CAPRI is designed to maintain strict target confidentiality, and imposes little delay on its publication.
More information about target submission is available at
the CAPRI site at http://bit.ly/vDLjWq and from the CAPRI
management team (joel.janin@u-psud.fr).
Notice for REFMAC Users
Depositors using REFMAC with TLS should make sure that the ATOM records contain full (and not residual) B factors before depositing an entry to the PDB.
When TLS is involved in refinement using a script, the keyword "tlsout addu" should be added to produce a coordinate file containing the full B factor. The latest REFMAC interface (in CCP4i) has a button "Add TLS contribution to XYZOUT" in the folder of TLS Parameters for adding this keyword.
If depositors have finished the final round of refinement without including "tlsout addu" keywords, and do not see ANISOU records in the coordinates, only the residual B factors have been generated in the coordinate file. The residual B factor needs to be updated to the full B factor in order to properly run validation checks.
If needed, the TLSANL page at deposit.rcsb.org/adit/REFMAC.html provides web form server and scripting information for
converting files.
|



 Depositing a protein-protein, protein-DNA, protein-RNA or protein-peptide complex? Consider submitting your structure as a
target for CAPRI after depositing your entry to the PDB.
Depositing a protein-protein, protein-DNA, protein-RNA or protein-peptide complex? Consider submitting your structure as a
target for CAPRI after depositing your entry to the PDB.