
for Structural Bioinformatics Protein Data Bank
Spring
2008
Number 37
|
sf-convert: A Format
Conversion Tool for Structure Factor Files sf-convert can then output the data formatted as mmCIF, MTZ, CNS, TNT, SHELX, EPMR, XTALVIEW, HKL2000, Dtrek, XSCALE, MULTAN, MAIN, or OTHER. sf-convert is available for download from sw-tools.pdb.org. EmDep2:
Deposit EM Maps at the MSD-EBI or RCSB PDB The EMDB contains experimentally determined 3D maps and associated experimental data and files. This improvement to EMDB services is the first product of a collaboration between the European Network of Excellence 3D-EM (www.3dem-noe.org) and the recently NIH-funded Partnership for a Unified Data Resource for CryoEM (emdatabank.org) This partnership is comprised of the European Bioinformatics Institute, the Research Collaboratory for Structural Bioinformatics at Rutgers, and the National Center for Macromolecular Imaging at Baylor College of Medicine. EMDB: www.ebi.ac.uk/msd-srv/docs/emdb EMD-1415.
2008
Deposition Statistics The entries were processed by wwPDB teams at the RCSB PDB, MSD-EBI, and PDBj. Of the structures deposited in the first quarter of 2008, 75% were deposited with a release status of "hold until publication"; 22.5% were released as soon as annotation of the entry was complete; and 2.5% were held until a particular date. 90% of these entries were determined by X-ray crystallographic methods; 9% were determined by NMR methods. 97% of these depositions were deposited with experimental data. As of February 1, 2008, the deposition of experimental data is required. During the same period of time, 1915 structures were released into the archive. Data
Processing Versioning Procedures Version 3.0 is the format used for files released as a result of the Remediation Project. Since August 1, 2007, all files processed and released into the archive have followed Version 3.1. When modifications have been made to files released prior to that date, they have been then re-released in Version 3.1. Version 3.1 differs from Version 3.0 in descriptions of the biological unit (REMARK 300/350), geometry (REMARK 500), atom/residues modeled as zero occupancy (REMARK 475/480), non-polymer residues with missing atoms (REMARK 610), and metal coordination (REMARK 620). Documentation describing the differences between these versions is available at www.wwpdb.org/docs.html. Since the beginning of March 2008, the REVDAT record indicates when a Version 3.0 file is re-released as Version 3.1 with the name "VERSN." For example, if the journal record has been updated in an entry that previously followed Version 3.0, the REVDAT would appear as: REVDAT
1 04-MAR-08 1ABC 1 JRNL VERSN There is no change to how depositors submit their files. Any required changes in nomenclature can be made automatically by the wwPDB during the annotation process. Documentation about file formats and the Remediation Project is available at www.wwpdb.org.
|
E-mail: info@rcsb.org • Web: www.pdb.org • FTP: ftp.wwpdb.org
The RCSB PDB is a member of the wwPDB (www.wwpdb.org)
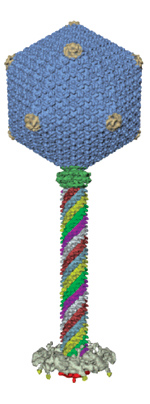 Electron
microscopy map data can now be deposited to the Electron Microscopy
Data Bank (EMDB) using the improved web-based tool EmDep2. EmDep2
is available from the existing deposition site at the MSD-EBI in
Europe and also from a new deposition site at the RCSB PDB in the
USA.
Electron
microscopy map data can now be deposited to the Electron Microscopy
Data Bank (EMDB) using the improved web-based tool EmDep2. EmDep2
is available from the existing deposition site at the MSD-EBI in
Europe and also from a new deposition site at the RCSB PDB in the
USA.