| EDUCATION
CORNER
Moving Pictures: Using Chimera to make
molecular multimedia for the classroom by Dr. Jeramia Ory, Kings
College

Jeramia Ory is an Assistant Professor
in the Department of Biology at King’s College in Wilkes-Barre,
Pennsylvania where he teaches Genetics, Biochemistry, and Systems
Biology. His training is in X-ray crystallography and NMR spectroscopy,
and he was previously a Biochemical Information Specialist at the RCSB
PDB. While at the RCSB PDB, he produced numerous images for this
newsletter, annual report and official documentation. The movies
and figures he uses for class can be viewed on his homepage (staff.kings.edu/
jeramiaory) in the “Multimedia” section, and are
free for educational use.
Getting students to grasp the link between 3D structure and biological
function is a necessary and challenging part of many undergraduate
courses. Structural information can help students “get it”
in a way that cannot be underestimated. As an example, numerous
students have told me how much easier it is to understand stereochemistry
when they can manipulate chemicals on a computer screen in 3D rather
than trying to work out wedge/hash 2D conventions. As the number
of structures in the PDB archive continues to grow, the challenge
lies not in finding structural information related to the topic
at hand (the Advanced Search on the RCSB PDB website is a great
resource), but in incorporating the information into lecture materials
and presentations without draining an instructor’s time or
resources. Fortunately for instructors, a number of free programs
that excel in molecular visualization and analysis are now available.
Instead of reviewing the myriad of programs out there, I will focus
the one I use to create multimedia presentations for my students–Chimera1.
CHIMERA is written
and maintained by the Computer Graphics Lab at the University of
California, San Francisco. It has a long history in molecular visualization,
having started as a program designed in 1980 for high-end graphics
workstations. What this means practically is that this research
group has been thinking about the needs of the molecular visualization
community for a long time. As modern desktop computing power has
grown, the visualization community has expanded from its original
base of X-ray crystallographers to educators and students as young
as high school. While no program can be all things to all people,
Chimera comes close. I have personally used it for hands on molecular
visualization workshops with groups ranging from high school students
to undergraduates with good results. Chimera has a few advantages
when compared to other packages out there.
COST. Chimera is free
for academic use. This is becoming less of a unique feature with
the rise of open source and free software, but it is still an important
consideration. After using Chimera for an exercise, I direct students
to the download page so they can use it on their home computers
if they wish to continue exploring. Just as important, Chimera is
available for every major operating system: Windows, Macintosh and
Linux (and more). Of course, as critics of free software are fond
of saying, “free software is only free if your time has no
value.” Luckily, the program is forgiving to new users, and
rewards time spent with it.
LEARNING CURVE. The last thing educators have is
time to waste. Chimera is a powerful analysis and visualization
tool and is written with scientists in mind, however, it is quite
easy to learn and new users can generally find their way around
the program in about an hour. I run a protein visualization exercise
in my undergraduate Biochemistry class that walks the students through
the basics of Chimera; they align myoglobin and hemoglobin and then
color the aligned residues by conservation (an example is shown
below).
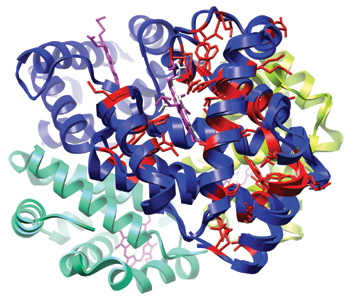
Myoglobin (2MM1) and hemoglobin
(4HHB) aligned; residues are colored by conservation
Most students complete the exercise in 50 minutes and find it useful
to be able to explore protein structure on their own. Should they
not finish, the fact that it is free means they can finish up at
the campus computer lab or at home. The program is well-documented
online (www.cgl.ucsf.edu/chimera)
and comes with tutorials for new users.
SUPPORT. The Chimera
community continues to grow as the program reaches different user
groups. There is an active mailing list of Chimera users that shares
ideas and problems, and is an excellent resource. Furthermore, the
developers of the program monitor the list, and have been known
to write modules for the program to deal with special users requests.
These requests have even been known to make it into future releases
as new features. One way or another, you can usually find someone
willing to help you make the figures or movies you want.
PRETTY PICTURES PRETTY FAST. Let’s face it,
you can stand in front of a lecture hall for an hour, waving your
arms, talking about the symmetrical relationships of hemoglobin’s
four subunits, or you can display some nicely rendered images and
a movie or two and get the same point across in five minutes. Say
what you will about today’s students, they respond to multimedia
and in some cases have grown to expect it. This is where Chimera
shines. Once learned, gorgeous images take just a few minutes to
set up and render. In my biochemistry class, I often make structural
figures rather than using the textbooks illustrations, or load Chimera
in class and walk students through the structure in 3D. This accomplishes
two things: 1) I get to highlight what it is I find important in
an RNA double helix, an enzyme active site, etc., and 2) it forces
me to learn the structural landscape of the molecules rather than
relying on the textbook. The rendering styles of Chimera are of
course a matter of taste, but frequent readers of the RCSB PDB’s
newsletter have already seen what Chimera can do; it is the “go
to” program among the staff and is used on many official publications.
So, now that you are convinced to give Chimera a try, how do we
make some movies? There are tutorials on the web site, but to start
with, you can try this simple set of commands. In your favorite
text editor (Notepad in Windows or TextEdit in Mac OS X), create
a file called “movie.cmd” and enter the following text:
movie record
roll y 1 360
wait 360
movie stop
movie encode
Save the file, start Chimera and load your molecule of interest.
Get it looking how you like, then open the file “movie.cmd”
(File… Open…) and watch it go. This script will rotate
the molecule about the y axis in 360 steps of 1 degree, then save
the movie. When it’s done you should have a file named “chimera_movie.mov”
that you can show to your students. If you would like to see some
of the movies I use in my Genetics and Biochemistry lectures, or
if you would like to use them in your own classes, please visit
my website.
- E.F. Pettersen, T.D. Goddard, C.C. Huang, G.S. Couch, D.M.
Greenblatt, E.C. Meng, and T.E. Ferrin (2004) UCSF Chimera–a
visualization system for exploratory research and analysis. J
Comput Chem. 25(13): 1605-12.
|


