Outreach and Education
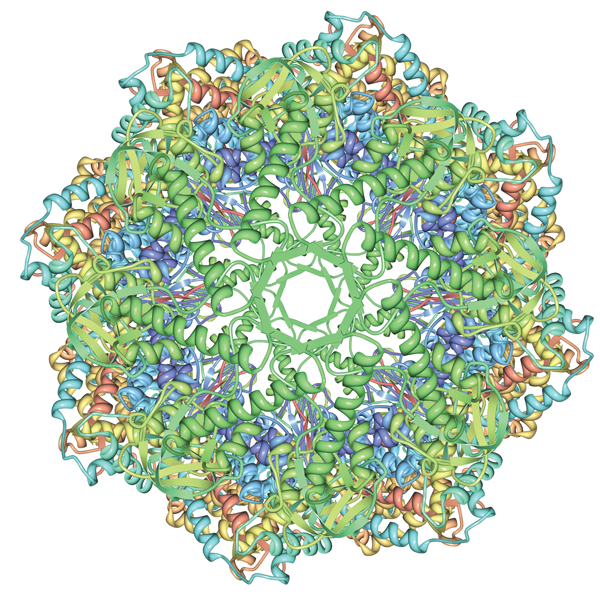
Image of 3izi created using Protein Workshop features.
High Resolution Images at the RCSB PDB
Create or download publication-quality pictures of biomacromolecules with the RCSB PDB.
Several interactive, Java-based tools2 can be used to visualize PDB data and create pictures. Protein Workshop offers easily customized views; Simple Viewer utilizes a quick ribbon display; and Ligand Explorer visualizes the interactions of bound ligands in protein and nucleic acids structures.
Each program can be used to create and save high-resolution images in JPEG, PNG, and TIFF formats. Using the Save Image dialog box from the File menu, users can specify the width and height of an image in pixels, inches, or millimeters.
From the educational PDB-101 site, high resolution TIFFs used to illustrate Molecule of the Month articles can be downloaded and used in presentations and publications (see citation and usage information).
PDB-101 posters, including The Structural Biology of HIV and Molecular Machinery: A Tour of the Protein Data Bank, can be saved as high or low resolution PDFs.
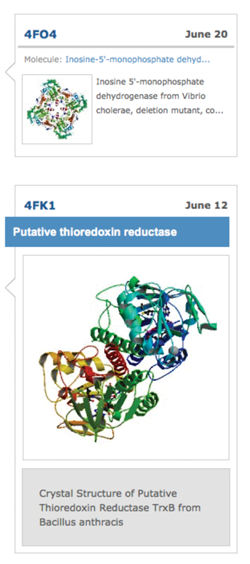
Author Profiles: Timeline Display of Structures
Author Profiles vertically display structures associated with a particular researcher. Profiles are now available for Structural Genomics centers. The timeline can be sorted by deposition date, and the first instance of a protein or complex in the author's profile is highlighted. Images are linked to the Structure Summary page for the entry.
To access this feature, select the link from the left hand RCSB PDB menu, or from the PDB-101 Features pull-down menu.
Image from the Author Profile for the Center for Structural Genomics of Infectious Diseases (CSGID)
Meeting News
ISMB and 3DSIG: The International Society for Computational Biology recently celebrated the 20th year of the ISMB–Intelligent Systems for Molecular Biology–conference (July 15-17, Long Beach, CA).
RCSB PDB presentations included Internal pseudo-symmetry in proteins (Andreas Prlić) and Efficient searching and mining of the RCSB Protein Data Bank (Peter Rose).
At the ISMB Special Interest Group meeting Bioinformatics Open Source Conference (BOSC), Andreas Prlić also presented How to use BioJava to calculate one billion protein structure alignments at the RCSB PDB website.
At the 3Dsig: Structural Bioinformatics & Computational Biophysics satellite meeting, John Westbrook discussed Format déjà vu: PDBx/mmCIF, the new data format for the wwPDB. Molecule of the Month author David Goodsell described Communicating and Interacting with the Molecular Cell (along with Arthur Olson).
Associate Director Phil Bourne was one of the 3Dsig Program Chairs.
ACA: From July 28–August 1, the RCSB PDB was at the 2012 Meeting of the American Crystallographic Association in Boston, MA. Attendees visited the exhibition booth to meet with members of the RCSB PDB and the PSI Structural Biology Knowledgebase and learned about new features such as RCSB PDB Mobile, Author Profiles, the wwPDB Common Tool for Deposition and Annotation, and much more. Lead annotator Jasmine Young presented an update on The Worldwide Protein Data Bank: Current Projects during the poster session. During the Structure-Guided Drug Discovery session, Data Management of Small Molecule Ligands, Antibiotics, and Peptide Inhibitors in the Protein Data Bank was described by RCSB PDB Director Helen Berman.
Protein Society: At the 26th Annual Symposium of The Protein Society (August 5–8, San Diego), Helen Berman was the 2012 Awardee of the Carl Brändén Award. The award, sponsored by the Rigaku Corporation, is given to an outstanding protein scientist who has also made exceptional contributions in the areas of education and/or service to the science. This award recognizes Dr. Berman's accomplishments toward enabling a freely available and uniform worldwide archive of 3D structural information for biomedical research and education. She presented Trendspotting from the Protein Data Bank as part of the Plenary Awards Session.
Poster Prizes Awarded
The RCSB PDB Poster Prize is awarded for the best student poster presentations at selected meetings. Recipients receive an educational book and a subscription to Science.

Sergei KalynychAt ACA, the award went to Sergei Kalynych for Crystallographic studies of closely related lipopolysaccharide O-antigen chain length regulators (Sergei Kalynych,1 Deqiang Yao,2 James Magee,1 Mirek Cygler;2
1McGill University, 2University of Saskatchewan)
An honorable mention was awarded to Rebecca Goldstein for A possible mechanism for the regulation of the PI-PLC from S. aureus (Rebecca Goldstein, Jiongjia Cheng, Mary Roberts; Boston College).
Many thanks to the judges: Barry Finzel (Chair, University of Minnesota), Patrick Loll (Drexel University), and David Rose (University of Waterloo), and to Marcia Colquhoun and the ACA.

Alan BarberAt ISMB, Alan Barber was recognized for Evolution of function in the alkaline phosphatase superfamily (Alan Barber,1 Jonathan Lassila,2 Helen Wiersma-Koch,2 Michael Hicks,1 Daniel Herschlag,2 Patricia Babbitt;1 1University of California, San Francisco, 2Stanford University).
Many thanks to Steven Leard, the International Society for Computational Biology, and the judges: Yana Bromberg (Chair, Rutgers), Hannah Carter (Johns Hopkins), Jeroen deRidder (Delft University of Technology), Iddo Friedberg (Miami University, Oxford, Ohio), Tatyana Goldberg (Technical University Munich, Germany), Ben Jelen (Rutgers), Hande Kucuk (University of Miami), Biao Li (Buck Institute, Novato), Magali Michaut (Netherlands Cancer Institute), Yanay Ofran (Bar Ilan University), Susanna Repo (EMBL-EBI), Venkata Satagopam (EMBL-Heidelberg), Andrea Schafferhans (Technical University Munich), Avner Schlessinger (UCSF), Stefan Senn (Rutgers), Janita Thusberg (Buck Institute, Novato), Tobias Wittkop (Buck Institute, Novato), Chengsheng Zhu (Rutgers).

Jonas LindholtAt the European Crystallographic Meeting (August 7-11, Bergen, Norway), Jonas Lindholt received the prize for Cardiotonic Steroids and the Na/K-ATPase (Jonas Lindholt, Linda Reinhard, Poul Nissen; Centre for Membrane Pumps in Cells and Disease (PUMPkin) and Department of Molecular Biology and Genetics, Aarhus University, Denmark).
Many thanks to Andreas Roodt (ECA Prize Committee and University of the Free State, South Africa), the European Crystallographic Association, and the judges: Udo Heineman (Max-Delbrück Center for Molecular Medicine, Berlin), Linda Shimon (Weizmann Institute of Science), and Ute Krengel (University of Oslo).
All poster prize awardees are listed on the RCSB PDB website.
wwPDB News
Future Funding of the BioMagResBank: White Paper and Community Discussions
A white paper on the future funding of the BMRB and related articles have been published in the September issue of Nature Structural & Molecular Biology:
Editorial: The BMRB matters (2012) NSMB 19: 853 doi:10.1038/nsmb.2387
Community Comments: In support of the BMRB (2012) NSMB 19: 854—860 doi:10.1038/nsmb.2371
From the BMRB: White Paper on Future Funding (doc)
This issue has also been discussed in the Nature, The Scientist, and Corante
The BMRB, housed at the University of Wisconsin—Madison, collects, annotates, archives, and disseminates the important spectral and quantitative data derived from NMR spectroscopic investigations of biological macromolecules and metabolites.
Current funding from the National Library of Medicine is set to expire in 2014. It appears critical that a funding plan be developed within the coming year.
